Overview Of Protista And Its Classification - Free Essay.
In the Systematics Lab, you can explore 5 different taxonomic classification schemes that biologists have used (from the traditional Linnaean scheme to the current 3-domain system). In this activity, you will learn how to use the Virtual Systematics Lab to identify the characteristics that various organisms share, and determine the relatedness of different taxa.
Based on a single posterior flagellum (opisthokont), flattened, nondiscoid mitochondrial cristae, a chitinous exoskeleton, storage of glycogen instead of starch, lack of chloroplasts, and the code UGA for tryptophan, not chain termination, in their mitochondria, Cavalier-Smith (54) suggests a common origin of the true fungi with animalia and choanoflagellate protozoa (fig. 1).
In cladistic classification, the contents of Protista are distributed among various supergroups (SAR, such as protozoa and some algae, Archaeplastida, such as land plants and some algae, Excavata, which are a group of unicellular organisms, and Opisthokonta, such as animals and fungi, etc.).
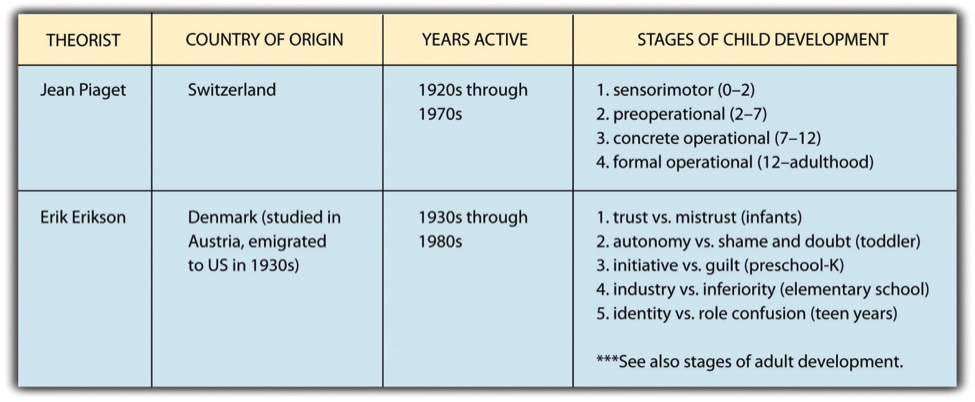
Protozoan, organism, usually single-celled and heterotrophic (using organic carbon as a source of energy), belonging to any of the major lineages of protists and, like most protists, typically microscopic. All protozoans are eukaryotes and therefore possess a “true,” or membrane-bound, nucleus.
The animal mode of nutrition is called heterotrophic because they get their food from other living organisms. Some animals eat only plants; they are called herbivores. Other animals eat only meat and are called carnivores. Animals that eat both plants and meat are called omnivores.
The classification proposed by Adl et al. (2005) on behalf of The Society established name stability as well as a synthesis of the overall structure of the classification of eukaryotes, based on the information available at that time, and after the upheaval introduced by molecular phylogenetic studies over the preceding two decades. Overall, the system proposed was conservative enough to.
Glomeromycota fungi contain several hundreds of nuclei that show a low level of intersporal (Tisserant et al., 2013) or internuclei polymorphism (Lin et al., 2014).However, the nuclear ribosomal encoding gene, in contrast to the rest of the genome, is highly diverse (Sanders et al., 1995; Lanfranco et al., 1999; Pawlowska and Taylor, 2004).The consequence is that several ribosomal sequences.